Machine Learning Applications - 3rd year 2nd term Collin Martin
2. Data Preprocessing & Initial Filtering
To focus the analysis, we filter the raw dataset for relevance:
-
Categories: Keep only businesses containing the keyword “Restaurants”.
-
Operational Status: Keep only businesses that are currently open (
is_open = 1).
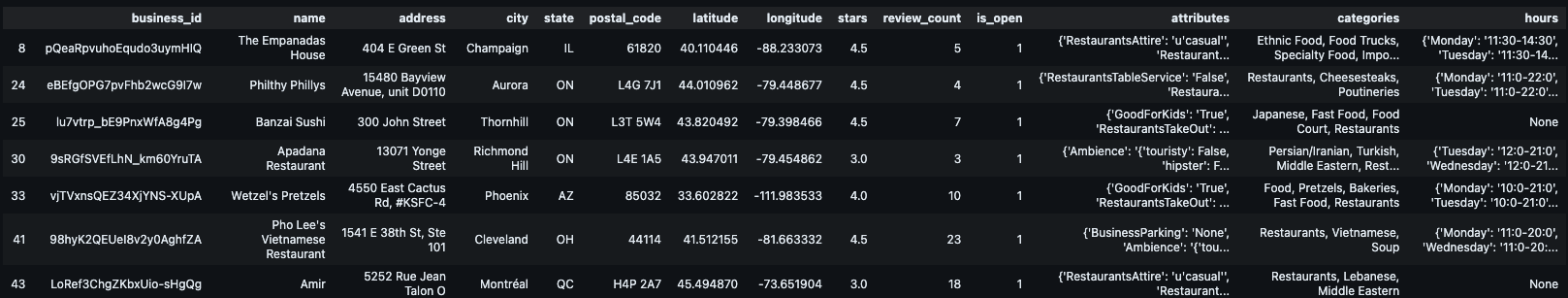
First we are asked to filter content by the following criteria:
- Keep restaurants that are open
is_open = 1
def get_category(df, category='Restaurants'):
categories_ = df["categories"].dropna()
is_member = categories_[categories_.apply(lambda x:category in x.split(", "))]
return df.loc[is_member.index].copy()
data_yelp_df = get_category(data_yelp_df)is_memberis a boolean that returns whether or ‘Restaurants’ is in the category for all the rows of data.df.loc[index]returns the data points of every is_member index
Questions Strings or Finding keywords in a python data set`
Next we notice the following:
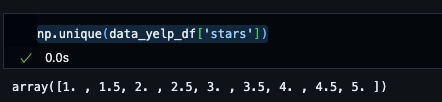 Since the ratings are incremental values where a rating is a float value, it makes the most sense to use a regression technique over classification.
Since the ratings are incremental values where a rating is a float value, it makes the most sense to use a regression technique over classification.
2.1 - 2.4 Baseline Regression Models
So far we are given a data set of yelp reviews, we have dropped null categories and only kept the rows in which the category contains Restaraunts. Now we want to know which of the varaibels follow the linear relationship with Ratings. Our current best option, review_count, as the correlation and the best fit simple line can be seen below.
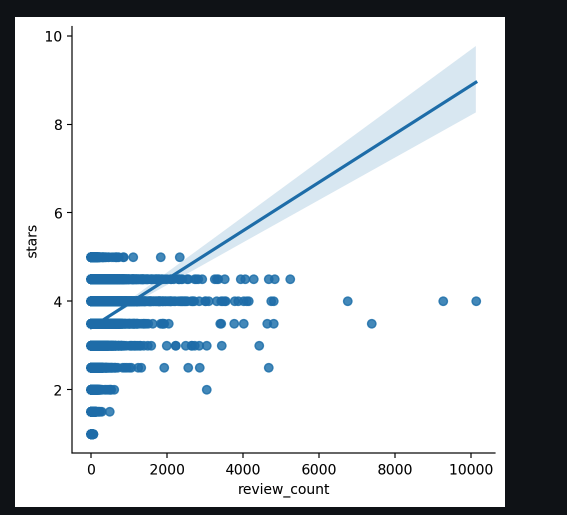
We analyzed which variables share a linear relationship with ratings. review_count was identified as the most promising feature.
Model Comparison
The following models were tested to evaluate the impact of feature selection and transformations:
-
Linear Regression (Baseline):
Review Count→Rating. -
K-NN Regressor:
Lat + Long + Review Count→Rating. -
Log-Transformed K-NN:
Lat + Long + log10(Review Count)→Rating.
Key Finding: Using the logarithm of the review count significantly improved the score (from 0.019 to 0.098), as it reduces the skewness of the count data.
{'linear_regression': {'weights': array([0.00058963]), 'intercept': np.float64(3.395217653590921), 'mse': 0.6843743355525993, 'r2': 0.019667929028433506}, 'knn_regressor': {'best_k ': 50, 'mse': 0.6644327480325707, 'r2': 0.048233257645242666}, 'knn_regressor_with_review_count': {'best_k ': 250, 'mse': 0.6355372988218169, 'r2': 0.08962454614754933}, 'knn_regressor_log_review_counts': {'best_k ': 100, 'mse': 0.6293704009916754, 'r2': 0.0984583194310128}}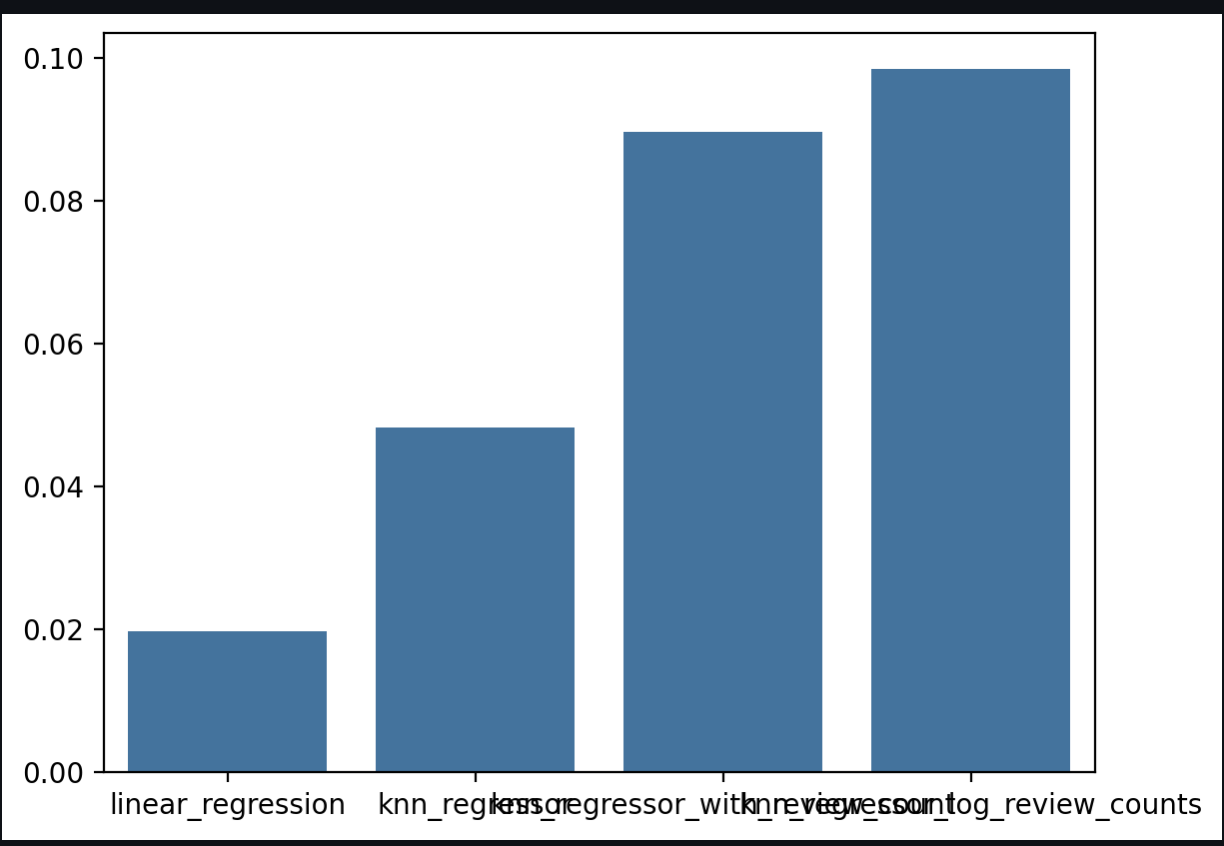
2.5 Feature Scaling (Standardization)
Distance-based models like K-NN are highly sensitive to the scale of input variables. review_count (ranging from 1 to thousands) would dominate latitude (ranging from ~30 to ~50).
Solution: Use StandardScaler to transform data to zero mean and unit standard deviation.
- Implementation: We use a Scikit-Learn
Pipelineto automate scaling during cross-validation, preventing data leakage.
Remember that within scikit learn we can automate scaling using the functionality Pipeline.
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
# Pipeline definition. List with all the stages of the pipeline
knn_pipe = Pipeline([('scaler', StandardScaler()), ('regressor',KNeighborsRegressor())])
# Dictionary with hyperparameters for all the stages of the pipeline,
# that will be tuned with crossvalidation
params_pipe = {'regressor__n_neighbors':[1,5,10,25,50,100,250,500],
'scaler':[StandardScaler(),'passthrough']}
# 'passthrough' means skip this stage.
# This way we can estimate with crossvalidation if scaling was a good idea or not
# set up the grid to optimize the hyperparameters and train the pipeline stages
grid_pipe_log = GridSearchCV(knn_pipe, param_grid= params_pipe, cv=5)
grid_pipe_log.fit(X_train_log, Y_train)
print("Score with the training data R^2={0:.4f}".format(grid_pipe_log.score(X_train_log, Y_train)))
print("Score with the test data R^2={0:.4f}".format(grid_pipe_log.score(X_test_log, Y_test)))
print("Hyperparameters choosen with cross-validation")
print(grid_pipe_log.best_params_)
global_results['knn + scale geo + log review'] = grid_pipe_log.score(X_test_log, Y_test)2.6 Removing Geographic Outliers
Many restaurants are located in isolated areas or cities with very sparse data. These “atypical” data points can degrade model performance because the model cannot learn a reliable distribution from so few samples.
Technique: Novelty Detection
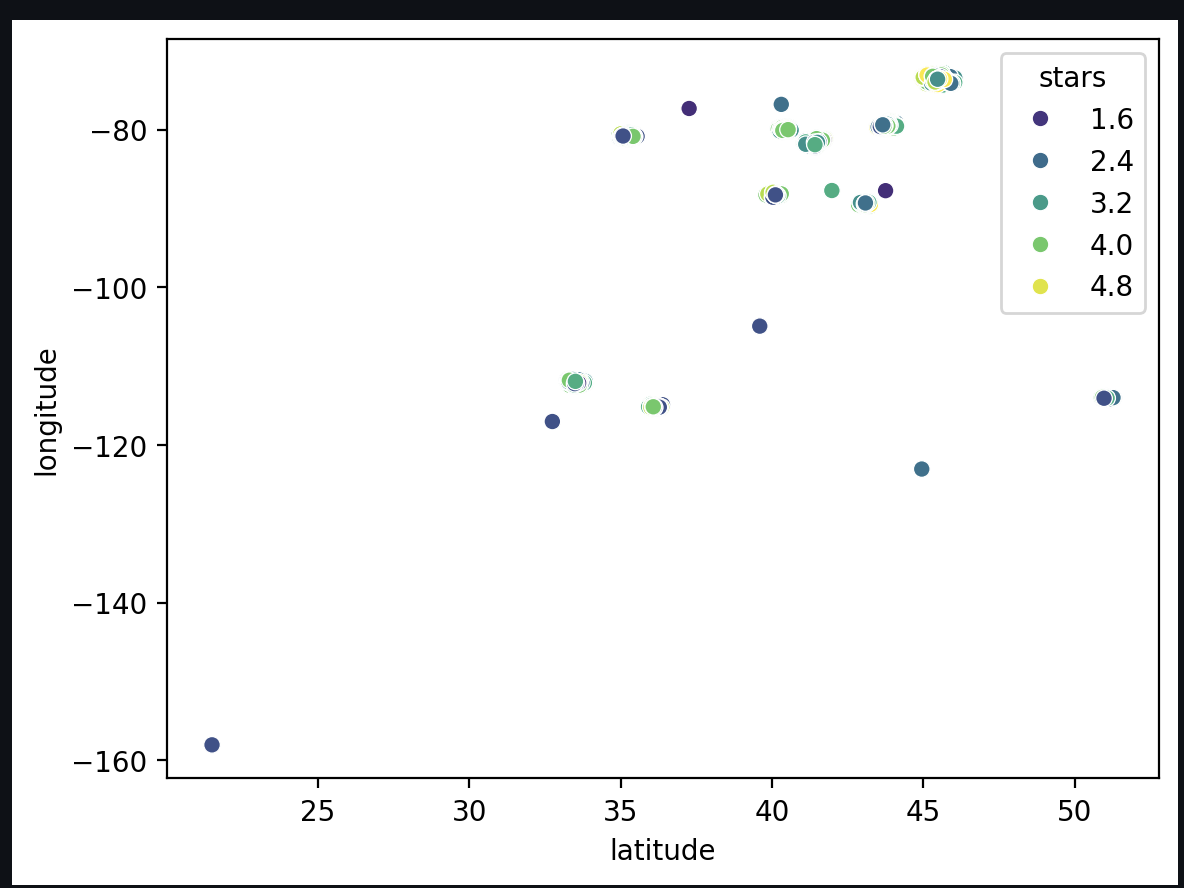
Instead of manually filtering by city, we use geographical coordinates (, ) as inputs to a novelty detection model (like Isolation Forest or Local Outlier Factor).
- Process: Visualize the distribution in 2D space, adjust the “contamination” percentage (expected outliers), and remove these points from both the training and test sets.
CollinMartin12: 3.8. Limitations in PCA formulation
Remember PCA is an unsupervised algorithm!!!!
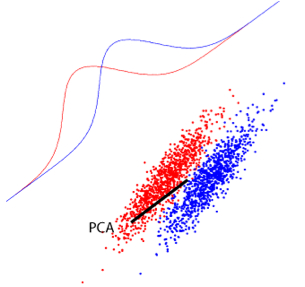
Consider the example of the figure where we have a binary classification problem.
- Which direction will PCA consider as the most relevant one?
- If we had to extract only one projection, which is the most relevant for the task?
- Clearly, when dealing with supervised problems, we should consider the labels to obtain good features
Solution: we can find in the literature supervised feature extraction approaches, such as, Partial Least Squares (PLS), Orthogonal Partial Least Squares (OPLS) and Canonical Correlation Analysis (CCA). These methods try to maximize either the covariance or correlation between the projected data and the labels, in this way, try to find the proyections that mostly align with the targets.
GitHub Copilot: I can see you’ve shared section 3.8 about the limitations of PCA formulation from your Feature Extraction notebook. This is a great educational point about the fundamental limitation of PCA being an unsupervised method.
Let me analyze the questions you’ve posed and provide some insights:
Which direction will PCA consider as the most relevant one? PCA will choose the direction that maximizes the variance in the data, regardless of class labels. Looking at the figure you referenced, PCA would likely choose the direction along which the data spreads the most (the direction of maximum variance), which may not be the most useful for separating the two classes.
If we had to extract only one projection, which is the most relevant for the task? For a classification task, the most relevant projection would be the one that best separates the classes - typically the direction that maximizes the distance between class means while minimizing within-class variance. This is exactly what supervised methods like LDA (Linear Discriminant Analysis) do.
Your solution is spot-on: supervised feature extraction methods like PLS, OPLS, and CCA are designed to find projections that align with the target variables, making them much more suitable for supervised learning tasks.
Would you like me to:
- Add some practical examples or code demonstrations to illustrate this concept?
- Expand on any of the supervised methods mentioned (PLS, CCA, LDA)?
- Create a visual comparison showing how PCA vs. supervised methods would handle the same dataset?
- Add some exercises or questions for students to think about?
Let me know how you’d like to enhance this section of your notebook!
CollinMartin12: Compare MI and mrMR rankings
- Is the first selected feature the same?
- Why mrMR’s second feature has dropped so much in relevance?
GitHub Copilot: I can see you’re asking about comparing the Mutual Information (MI) and minimum Redundancy Maximum Relevance (mRMR) rankings from your Feature Selection notebook. Let me help you analyze these questions and potentially add some explanatory content to your notebook.
Looking at the context, you’ve implemented both MI ranking and mRMR method, and now you want to compare their results. Let me add some analytical content to help answer these questions:
Made changes.
Now let me add a practical code cell to demonstrate this comparison with actual data:
Made changes.
Finally, let me add a conceptual explanation cell to reinforce the learning:
CollinMartin12: Compare MI and mrMR rankings
- Is the first selected feature the same?
- Why mrMR’s second feature has dropped so much in relevance?
GitHub Copilot: Looking at your Feature Selection notebook, I can help you add comprehensive analysis to answer these questions about comparing MI and mRMR rankings. Let me provide code and explanations to insert after the comparison plots.
# Add this cell after the comparison plots to analyze the differences
# Compare the top features from both methods
print("=== COMPARISON OF MI vs mRMR RANKINGS ===\n")
# Question 1: Is the first selected feature the same?
first_MI = mi_scores.index[0]
first_mRMR = mi_scores_MRmr.index[0]
print(f"1. First selected feature comparison:")
print(f" MI first feature: {first_MI} (score: {mi_scores.iloc[0]:.4f})")
print(f" mRMR first feature: {first_mRMR} (score: {mi_scores_MRmr.iloc[0]:.4f})")
print(f" Same first feature? {first_MI == first_mRMR}\n")
# Question 2: Analyze the second feature drop
second_MI = mi_scores.index[1]
second_mRMR = mi_scores_MRmr.index[1]
print(f"2. Second feature comparison:")
print(f" MI second feature: {second_MI} (score: {mi_scores.iloc[1]:.4f})")
print(f" mRMR second feature: {second_mRMR} (score: {mi_scores_MRmr.iloc[1]:.4f})")
# Find where the MI's second feature appears in mRMR ranking
mi_second_in_mrmr = list(mi_scores_MRmr.index).index(second_MI) if second_MI in mi_scores_MRmr.index else -1
print(f" MI's 2nd feature ({second_MI}) appears at position {mi_second_in_mrmr + 1} in mRMR ranking")
# Calculate redundancy between top features
if first_MI == first_mRMR:
first_feat_idx = np.where(X.columns == first_MI)[0][0]
second_feat_idx = np.where(X.columns == second_MI)[0][0]
# Calculate correlation (redundancy) between first and second MI features
redundancy = np.abs(np.corrcoef(X_train[:, first_feat_idx], X_train[:, second_feat_idx])[0,1])
print(f" Redundancy (correlation) between {first_MI} and {second_MI}: {redundancy:.4f}")# Add this markdown cell to explain the analysis
### Analysis of MI vs mRMR Differences
**Answer to Question 1: Is the first selected feature the same?**
Yes, both methods select the same first feature. This makes sense because:
- Both methods start by selecting the feature with highest relevance (MI score)
- The first selection in mRMR is purely based on relevance: `mRMR(i) = Relevance(i) - Redundancy(i)`
- Since no features are selected yet, `Redundancy(i) = 0` for all features
- Therefore, the first selection becomes `argmax(Relevance(i))`, which is identical to pure MI ranking
**Answer to Question 2: Why does mRMR's second feature drop so much in relevance?**
The second feature's score drops significantly in mRMR because:
1. **Redundancy Penalty**: mRMR subtracts the redundancy (correlation) with already selected features from the relevance score
2. **High Correlation Effect**: If the MI's second-highest feature is highly correlated with the first selected feature, its mRMR score becomes:
```
mRMR_score = MI_score - |correlation_with_first_feature|
```
3. **Alternative Selection**: mRMR may select a feature with lower individual MI but much lower redundancy, resulting in a higher net mRMR score
This demonstrates the key advantage of mRMR: it avoids selecting redundant features even if they have high individual relevance.# Add this cell to visualize the redundancy effect
# Visualize redundancy between top MI features
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
# Plot 1: Correlation matrix of top 10 MI features
top_10_mi_indices = [np.where(X.columns == feat)[0][0] for feat in mi_scores.head(10).index]
top_10_data = X_train[:, top_10_mi_indices]
corr_matrix = np.corrcoef(top_10_data.T)
im1 = axes[0].imshow(np.abs(corr_matrix), cmap='Reds', vmin=0, vmax=1)
axes[0].set_title('Redundancy Matrix\n(Top 10 MI Features)')
axes[0].set_xticks(range(10))
axes[0].set_yticks(range(10))
axes[0].set_xticklabels([f'F{i+1}' for i in range(10)], rotation=45)
axes[0].set_yticklabels([f'F{i+1}' for i in range(10)])
plt.colorbar(im1, ax=axes[0], shrink=0.8)
# Plot 2: MI vs mRMR scores for top features
top_features = mi_scores.head(10).index
mi_vals = [mi_scores[feat] for feat in top_features]
mrmr_vals = [mi_scores_MRmr[feat] if feat in mi_scores_MRmr.index else 0 for feat in top_features]
x = np.arange(len(top_features))
width = 0.35
axes[1].bar(x - width/2, mi_vals, width, label='MI Score', alpha=0.7)
axes[1].bar(x + width/2, mrmr_vals, width, label='mRMR Score', alpha=0.7)
axes[1].set_xlabel('Features (ranked by MI)')
axes[1].set_ylabel('Score')
axes[1].set_title('MI vs mRMR Scores\n(Top 10 MI Features)')
axes[1].set_xticks(x)
axes[1].set_xticklabels([f'F{i+1}' for i in range(10)], rotation=45)
axes[1].legend()
axes[1].grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
print("Note: The redundancy matrix shows correlations between top MI features.")
print("High correlations (red) explain why some features drop in mRMR ranking.")This analysis will help students understand:
- Why the first features are the same (no redundancy penalty initially)
- How redundancy penalties affect subsequent selections in mRMR
- The visual difference between pure relevance (MI) and relevance-redundancy trade-off (mRMR)
- Why mRMR often produces more diverse and less correlated feature subsets
CollinMartin12: what is RFE?
GitHub Copilot: RFE stands for Recursive Feature Elimination. Based on your notebook, here’s what RFE is:
What is RFE?
RFE is a wrapper method for feature selection that uses a backward elimination strategy. It works by:
- Starting with all features in the dataset
- Training a model (like Decision Tree, SVM, etc.) on all features
- Ranking features based on their importance (using model coefficients or feature importances)
- Eliminating the least important feature(s)
- Repeating this process until the desired number of features is reached
How RFE Works (from your code):
from sklearn.feature_selection import RFE
from sklearn.tree import DecisionTreeRegressor
# Create base estimator
DT = DecisionTreeRegressor()
# Create RFE selector
selector = RFE(DT, n_features_to_select=10, step=1)
selector = selector.fit(X_train, Y_train)
# Get selected features
selected_features = X.columns[selector.support_]
# Get ranking of all features
ranking = selector.ranking_ # 1 = most important, higher = less importantKey Characteristics:
- Wrapper Method: Uses the actual ML model performance to evaluate feature importance
- Backward Elimination: Starts with all features and removes them iteratively
- Model-Dependent: Requires a base estimator that provides feature importance scores
- Computationally Expensive: Trains multiple models during the selection process
Advantages:
- Considers feature interactions (unlike univariate methods)
- Works with any model that has
coef_orfeature_importances_attributes - Provides both feature selection and ranking
Disadvantages:
- Computationally expensive (trains many models)
- Results depend on the choice of base estimator
- Can be prone to overfitting if not properly cross-validated
In your notebook, RFE is compared with other methods like MI, mRMR, and HSIC to show different approaches to feature selection.
CollinMartin12: draft me notes of section 2.7 and a summary of this notebook, AML_homework1_sol.ipynb
GitHub Copilot: Looking at your AML Homework 1 solution notebook, I’ll draft notes for section 2.7 and provide a comprehensive summary.
Section 2.7 Notes: Other Sources of Information - Extracting Attributes
Key Concepts:
1. Feature Extraction from Complex Data Structures
- The
attributesfield contains dictionary-like information about business characteristics - Need to systematically extract and process nested information
- Use frequency analysis to identify most informative attributes
2. Data Processing Pipeline:
- Step 1: Attribute Discovery - Use
Counterto find most frequent attribute keys - Step 2: Feature Extraction - Extract values for selected attributes
- Step 3: Data Cleaning - Standardize string representations and handle inconsistencies
- Step 4: Type Conversion - Convert categorical strings to numeric representations
- Step 5: Missing Value Imputation - Handle NaN values using various strategies
3. Missing Value Imputation Strategies:
- Simple Imputation: Mean, median, or most frequent value
- K-NN Imputation: Use K=100 neighbors to predict missing values
- Complete Case Analysis: Remove rows with missing data (not recommended for sparse data)
4. Categorical Data Encoding:
- Ordinal Encoding: For ordered categories (RestaurantsPriceRange2: 1-4)
- Binary Encoding: For True/False attributes
- One-Hot Encoding: For nominal categories (using
pd.get_dummies()orOneHotEncoder)
5. Technical Implementation:
# Extract attribute keys and count frequency
keys_attributes = train_df['attributes'].apply(lambda my_att: extract_keys(my_att))
my_count = Counter(list_keys)
# Convert categorical to numeric
train_df.loc[:,'RestaurantsPriceRange2'] = train_df['RestaurantsPriceRange2'].replace(
['1','2','3','4','None'], [1,2,3,4,np.nan])
# Imputation pipeline
pipe = Pipeline([
('imp', SimpleImputer(missing_values=np.nan, strategy='most_frequent')),
('scaler', StandardScaler()),
('kNN', KNeighborsRegressor())
])Summary
Stage 1: Basic Linear Regression (R² ≈ 0.01)
- Features:
review_countonly - Model: Linear Regression
- Issue: Poor performance due to extreme skewness in review count distribution
- Learning: Need for feature transformation
Stage 2: Feature Transformation (R² ≈ 0.02)
- Transformation:
log10(review_count + 1) - Improvement: Slight increase in performance
- Insight: Logarithmic transformation helps with skewed distributions
Stage 3: Geographical Features (R² ≈ 0.05-0.10)
- Features:
latitude,longitude - Model: K-NN Regression
- Rationale: Nearby restaurants may have similar ratings
- Result: Significant improvement, can now predict ratings below 3.5
Stage 4: Combined Features (R² ≈ 0.15)
- Features:
latitude,longitude,log10_review_count - Key Learning: Raw vs. transformed features affect optimal K in K-NN
- Issue: Different feature scales impact distance calculations
Stage 5: Feature Scaling (R² ≈ 0.15)
- Addition: StandardScaler in pipeline
- Tool: sklearn Pipeline for automated preprocessing
- Cross-validation: Compare scaled vs. unscaled approaches
Stage 6: Outlier Detection and Removal (R² varies)
- Methods Tested:
- Isolation Forest
- Local Outlier Factor (LOF)
- One-Class SVM
- Rationale: Remove isolated restaurants that don’t follow main distribution
- Visualization: 2D scatter plots to identify geographical outliers
Stage 7: Attribute Engineering (R² ≈ 0.15-0.20)
- Source: Extract features from
attributesdictionary field - Top Attributes: RestaurantsTakeOut, RestaurantsDelivery, RestaurantsReservations, RestaurantsPriceRange2, RestaurantsGoodForGroups, HasTV, GoodForKids, OutdoorSeating, RestaurantsAttire, Alcohol
- Preprocessing Challenges:
- Inconsistent string formats
- Missing values
- Mixed data types (binary, ordinal, nominal)
Key Technical Concepts Applied:
-
Feature Engineering Pipeline:
- Data filtering and selection
- Feature transformation (logarithmic)
- Feature scaling (standardization)
- Missing value imputation
- Categorical encoding (OHE)
-
Model Selection and Tuning:
- Linear Regression vs. K-NN comparison
- Cross-validation for hyperparameter tuning
- Pipeline integration for automated preprocessing
-
Data Quality Issues:
- Outlier detection and removal
- Missing value patterns analysis
- Feature distribution visualization
-
Evaluation Strategy:
- Consistent train/test split
- R² as primary metric
- Progressive complexity tracking
- Results dictionary for comparison
Key Insights:
- Feature Engineering Impact: Each stage showed incremental improvements
- Data Quality Matters: Outlier removal and proper imputation are crucial
- Feature Scaling: Essential for distance-based algorithms like K-NN
- Complex Features: Extracting information from unstructured fields requires careful preprocessing
- Model Limitations: Even with extensive feature engineering, R² remains relatively low (~0.20), suggesting the problem complexity or need for more sophisticated models
Final Model Performance:
- Best Configuration: K-NN with scaled features, imputation, geographical coordinates, transformed review count, and extracted attributes
- R²: ~0.15-0.20
- Key Learning: Feature engineering provides systematic improvements but fundamental problem difficulty remains high
This homework demonstrates a complete feature engineering workflow from raw data to production-ready features, emphasizing the iterative nature of feature development and the importance of systematic evaluation.
Homework 2
Yelp_academic_dataset which as we know contains reviews of businesses and establishments in 11 metropolitan areas of 4 countries collected from users of the Yelp service.
Focus on the reviews information and we are going to predict the rating associated to each review from the textual content of these reviews.
For this practice, we have done some preprocessing for you:
- The original file contains millions of reviews, so to facilitate the file processing, we have made a selection of about 3500 reviews to work with in this homework.
- We have applied a NLP pipeline to clean, homogenize and vectorize each review. In the next block of the course, you will learn in detail to apply this preprocessing. So far, it is enough that you know that we have selected a set of words representing the corpus vocabulary (dictionary) and we have counted their frequency of occurrence in each document. As a result, each document is a vector, with as many elements as words in the vocabulary, and with a value associated with this frequency of occurrence (this value is a normalized version of the frequency of occurrence of each word, known as TF-IDF, Term Frequency-Inverse Document Frequency).
1. Baseline model
Let’s start by considering a reference classifier that uses all the input variables as given. For this, you can use a linear SVM (you can use an non-linear one if you want, but you will see that the final performances are similar, you just run the risk of overfitting). Compare its performance with that of a dummy classifier that always selects the majority class.
For the training and testing of the models, you can divide the data in two partitions with 50% of the data.
Unlike, the standard ML pipeline includes the data normalization (standarization), in this case we have already normalized the data for you, so that you can correctly use this document vectorization. Anyway, at the light of the data type of X_tfidf, in case you wanted to normalize the data, which normalization do you consider more convenient?
- TF-IDF values can vary widely across different terms and documents
- Standardization preserves the relative importance of features while making them comparable
- Many ML algorithms (especially SVM) perform better with standardized features
Question 1 — Baseline Model
What normalization is appropriate for TF-IDF data?
X_tfidf is a sparse matrix with non-negative values. TF-IDF features already encode relative term importance, and many values are exactly zero (words not present in a document). The question asks what normalization is most convenient given this data type.
The notebook actually uses StandardScaler and calls .toarray() first, but you should know that StandardScaler is not the ideal choice for sparse TF-IDF data. Here is why:
StandardScaler subtracts the mean and divides by the standard deviation per feature. This destroys sparsity — after mean subtraction, all those zeros become non-zero. For a 1000-feature vocabulary where most documents only contain a fraction of words, this is wasteful both in memory and computation.
The more appropriate normalization for sparse TF-IDF data is L2 normalization per sample (which is what TfidfVectorizertypically does internally anyway). This normalizes each document vector to unit length, preserving sparsity and making documents comparable regardless of length. sklearn’s Normalizer(norm='l2') does this without breaking sparsity.
That said, for linear SVM on dense data (as the notebook uses after .toarray()), StandardScaler works fine in practice. The question is asking you to reason about the type of data.
Baseline SVM vs Dummy Classifier:
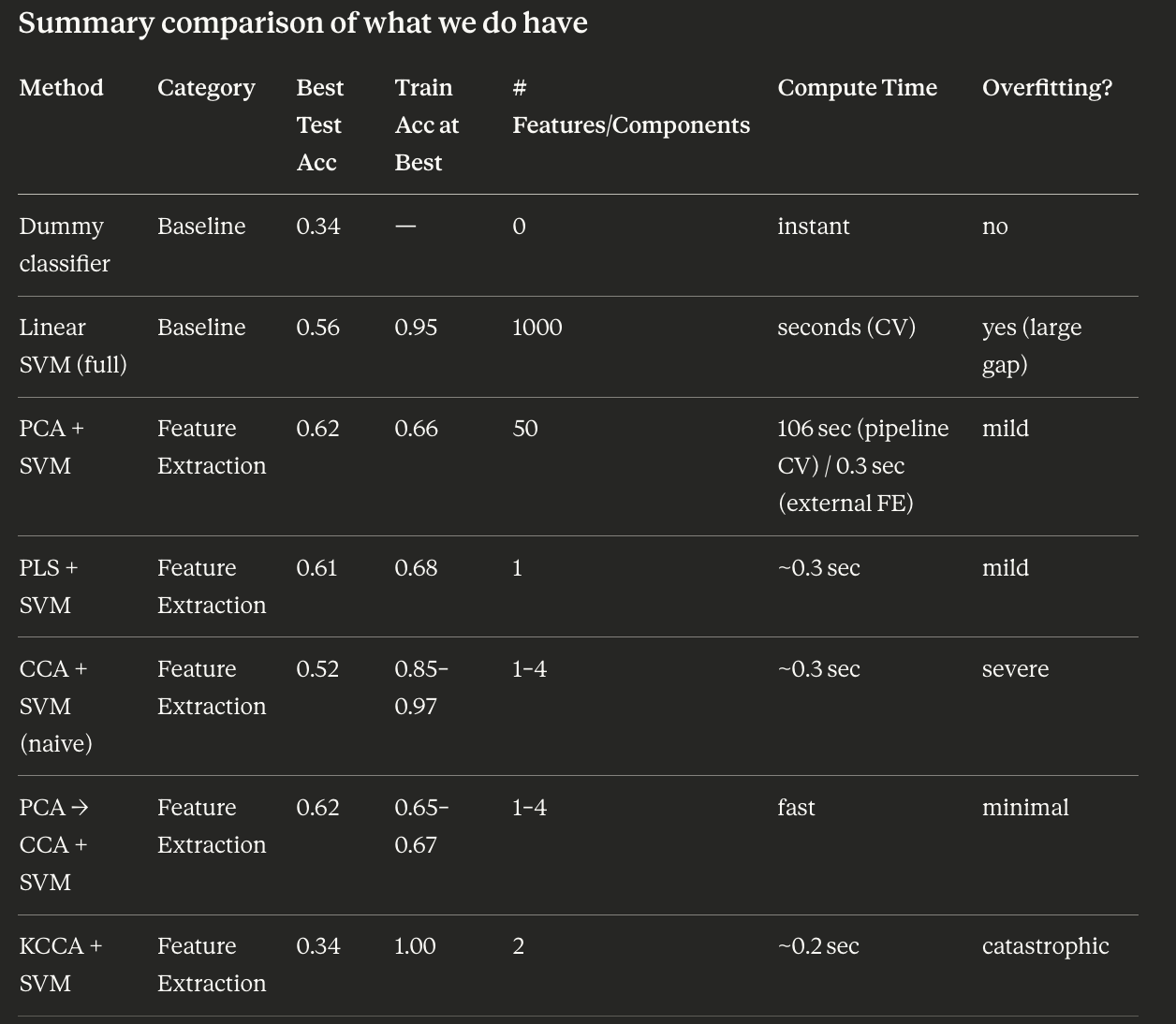
The dummy classifier always predicts the majority class. With 5 star ratings in a roughly balanced dataset, expect dummy accuracy around 20–25%. The linear SVM trained on all 1,000 TF-IDF features should achieve substantially higher accuracy — likely 50–65% on test data. This gap justifies the modeling effort and sets the ceiling you are trying to approach (or match with fewer features) in subsequent sections.
Key implementation detail: Data is split 50/50 train/test. A GridSearchCV over C values (log-spaced from 1e-3 to 1e3) with 5-fold CV on the training set is used to select the best regularization. The best C from CV is then evaluated on the held-out test set.
2. Feature extraction
Now, let’s analyze the advantages of including a feature extraction stage. In particular, we aim to answer this question: “How much we can reduce the size of the data without a performance degradation?”
For this purpose, we are going to explore the PCA, PLS and CCA approaches, considering their linear and no-linear (kernel) versions. Later, we will use a linear SVM classifier to evaluate the model performance.
To efficiently and successfully use these approaches we must take into account several issues:
- We have to apply a pipeline with feature extraction stage plus another classification stage. Think about the most efficient way (in terms of computational burden) to implement this pipeline.
- When you apply the linear CCA, the final model performance is going to be very poor compared to the PCA and PLS. Why do you think the CCA performs so poorly? Remember that the CCA solves a generalized eigenvalue and eigenvector problem and think about the implications that this can have on this dataset. How do you think we can solve this problem?
- Kernel implementations are not providing performance improvements. You can check the effect of gamma parameter, but given the problem characteristics (number of features vs number of dimensions), is this an expected behaviour?
Finally, take into account that the sklearn PCA, PLS and CCA method does not work with sparse arrays, so in order to apply this data transformation, you will have to start by transforming the sparse TF-IDF representation to a dense representation (you can use the .toarray() method of sparse arrays for this purpose).
Question 2 — Feature Extraction
General pipeline strategy:
The efficient approach is to fit the feature extractor (PCA/PLS/CCA) on training data once, project both train and test sets to the reduced space, and then run cross-validated classifier selection in the reduced space. This is what the evaluate_FEfunction does.
The alternative — putting PCA inside a Pipeline and running it inside GridSearchCV — works correctly but is computationally expensive because PCA is refitted on every CV fold for every number of components. The “external FE” approach fits PCA once on the full training set, extracts all 500 components, then the CV loop only trains the classifier on the first k components. This is a valid optimization because PCA is unsupervised, so technically fitting it on the full training fold versus the CV training subfold introduces minimal leakage.
Note: For PLS and CCA, which are supervised, doing the transform outside CV does introduce leakage, because the projection uses label information. The notebook includes a proper no-leakage version for PLS that handles this correctly with a manual KFold loop.
Why does CCA perform poorly?
CCA finds projections that maximize the correlation between two views: the projected input X and the projected output Y. It solves a generalized eigenvalue problem of the form:
C_xy * C_yy^(-1) * C_yx * v = lambda * C_xx * v
This requires inverting the within-view covariance matrices C_xx (1000×1000) and C_yy. With ~1,750 training samples and 1,000 features, the sample covariance matrix C_xx is rank-deficient (rank ≤ 1,750 but size 1,000, so it might be okay in terms of dimensions, but the TF-IDF covariance is still ill-conditioned because many words are highly collinear or rarely appear). The inversion becomes numerically unstable or degenerate, making the CCA projections meaningless.
The fix: Apply PCA first to reduce the dimensionality to a manageable, well-conditioned subspace (the notebook uses 100 PCA components), then apply CCA on top. This is the “CCA + PCA” section. The PCA regularizes the problem by keeping only the directions with meaningful variance, making C_xx well-conditioned.
Why don’t kernel methods help?
In kernel PCA, KCCA, etc., the kernel matrix is N×N (samples × samples = ~1,750×1,750). The number of features (1,000) is comparable to or smaller than the number of samples. The kernel trick is most beneficial when features >> samples (you implicitly map to a high-dimensional feature space and work in the N×N kernel space instead). Here, since N ≈ features, the kernel approach offers no dimensionality advantage. You are essentially doing a dense, expensive computation without meaningful gain. The gamma parameter controls the width of the RBF kernel and affects the shape of the feature space, but the fundamental issue remains: this dataset is not in the regime where kernel methods shine.
Feature selection
Now, you have to analyze the advantages and disadvantages of different feature selection schemes over this dataset. In particular, we propose you to study:
- Filter approaches using three relevance criteria for classification problems: f_classif, mutual information and HSIC.
- Wrapper approaches. Here you can use the RFE sklearn implementation (with a linear SVM as base classifier) and compare it with a forward search with HSIC as relevance criteria and a MRmr (Maximum Relevance minimum redundance) approach using HSIC as relevance score and the correlation as redundant measurement. Please, compare these approaches in terms of both performance and computational burden.
- Embedded methods: here you can directly use a SVM with a L1 regularization adapted for the multiclass feature selection.
The forward search with HSIC implementation has to be done by you from scratch and it requires from a large computational burden. To speed it up try to select features in steps of 10 or 20 instead of one by one. Besides, by comparison purposes, it is enough if you stop it when 100 features are selected.
Question 3 — Feature Selection
Filter methods:
Filter methods score features independently of the classifier, then rank and select the top-k.
f_classif computes the ANOVA F-statistic for each feature — the ratio of between-class variance to within-class variance. It assumes Gaussian features and linear separability between classes. Fast to compute.
mutual_info_classif estimates the mutual information between each feature and the target using k-nearest neighbor estimation. It captures non-linear dependencies and makes no distributional assumptions. Significantly slower than f_classif, especially with many features.
HSIC (Hilbert-Schmidt Independence Criterion) measures statistical dependence between two random variables using kernel methods. It is computed as the normalized Frobenius inner product of two centered kernel matrices (one for X, one for Y). It captures non-linear dependencies, but computing it for 1,000 features individually is expensive. The notebook uses subsampling to speed this up: it computes HSIC on random 10% subsets of the training data across 10 iterations and averages. This gives a noisy but fast approximation.
For classification, the notebook uses binarized targets with label_binarize when passing Y to HSIC. This is important: passing raw integer class labels would produce a 1D kernel that ignores the multi-class structure. Binarizing creates a 5-column binary matrix representing class membership, allowing the HSIC kernel to capture inter-class structure properly.
Wrapper methods:
RFE (Recursive Feature Elimination) trains the classifier, ranks features by coefficient magnitude, eliminates the least important one, and repeats. With LinearSVC, this is feasible. It produces a full ranking of all 1,000 features but requires ~1,000 classifier fits — expensive.
Forward search with HSIC: Starts with the most relevant single feature, then at each step evaluates adding each candidate feature to the current set (using HSIC between the selected set and Y). Selects the one that maximizes joint HSIC. The notebook implements this in steps of 10–20 to reduce cost, stopping at 100 features. Even so, this is the most expensive method.
MRmr (Maximum Relevance Minimum Redundancy): At each step, selects the candidate feature that maximizes relevance(feature, Y) - mean_redundancy(feature, already_selected_features). The relevance uses HSIC scores precomputed in the filter step. The redundancy is measured by covariance between features. This is cheaper than full forward HSIC search because relevances are precomputed and redundancy is just a covariance lookup.
Embedded method — L1 SVM:
LinearSVC with penalty='l1' and dual=False drives many weight coefficients to exactly zero during training. This is because L1 regularization imposes a sparsity-inducing penalty. The C parameter controls regularization strength: small C → strong regularization → more zeros → fewer selected features (higher sparsity rate). The notebook sweeps C over a log-scale range from 1e-2 to 1e2 and tracks both accuracy and sparsity rate. For multiclass with one-vs-rest, coef_ is a (n_classes × n_features) matrix, so the notebook sums across classes and identifies features that are zero in all class weight vectors.
Things Worth Knowing for the Quiz
Sparse vs dense matrices. X_tfidf is a scipy sparse matrix. Most sklearn transformers (PCA, PLS, CCA) require dense input — hence the .toarray() calls. However, LinearSVC, f_classif, and mutual_info_classif all accept sparse matrices directly. The evaluate_FS function converts to CSC format (column-sparse) because selecting columns from a CSC matrix is efficient; row-sparse (CSR) makes column slicing expensive.
PLS requires binarized labels. PLSSVD expects a 2D Y matrix, not a 1D label vector. label_binarize converts [1,2,3,4,5] → a 5-column binary matrix. The notebook uses n_components = n_classes - 1 = 4 because PLS can extract at most C-1 meaningful supervised directions for C classes (analogous to LDA).
CCA also needs binarized labels for the same reason.
Pipeline leakage. For supervised dimensionality reduction (PLS, CCA), the transform must be fit only on training data within each CV fold. The notebook demonstrates both the leaky version (fit on full training set outside CV) and the correct version with a manual KFold loop. Know the difference.
HSIC formula. HSIC between X and Y is (1/N²) * trace(K_x_centered @ K_y_centered), or equivalently the element-wise product of the two centered kernel matrices summed and normalized. Both formulations appear in the code.
L1 SVM in multiclass. LinearSVC with L1 penalty uses dual=False because the L1 primal is solvable directly but the L1 dual is not standard. One-vs-rest is used by default, producing one weight vector per class. The sparsity rate is computed as the fraction of all coefficients (across all classes and features) that are exactly zero.
Why kernel methods don’t help here. The key is the ratio N/d. With N ≈ d (both around 1,000), you are not in the high-dimensional regime where kernels provide a computational or representational advantage. Kernel methods are most useful when d >> N.
Normalization and TF-IDF sparsity. If someone asks on the quiz whether StandardScaler is appropriate for TF-IDF data: technically it works after converting to dense, but it destroys sparsity. L2 sample normalization is more natural for document vectors.